|
Instructions for the numerical files. 1) The codes The numerical analysis was done by the authors using Mathematica and Fortran programs. The Mathematica codes are used to produce all the figures of the paper and include a code that combine all the observational probes the authors used (it allows the user to choose the ones he wants) and performs a chi-square fit to the user’s model, it produces 1 sigma and 2 sigma confidence contours and the best fit form of the equation of state along with the 1sigma error region. For detailed instructions see the Mathematica codes section. Also the authors supply their modified version of CAMB. It can calculate the power spectrum of models that have an evolving equation of state parameter of the form w(a)=w0+w1(1-a), where a is the scale factor and w0,w1 are two constants. The code can also calculate the power spectrum for a grid of models and a set of pairs of (w0,w1) values that are input from a file and more importantly the likelihood of that model by using the WMAP 3 likelihood code. For detailed instructions see the Fortran codes section and the
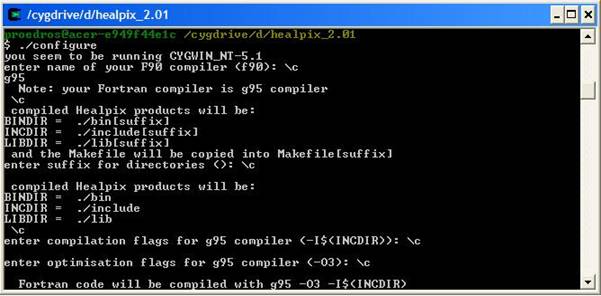
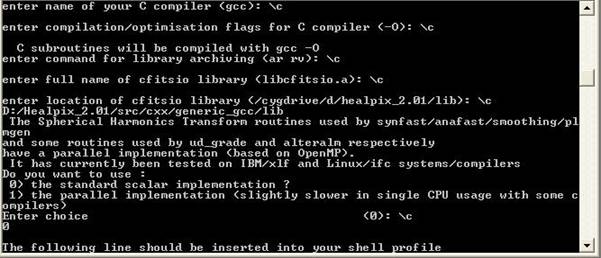
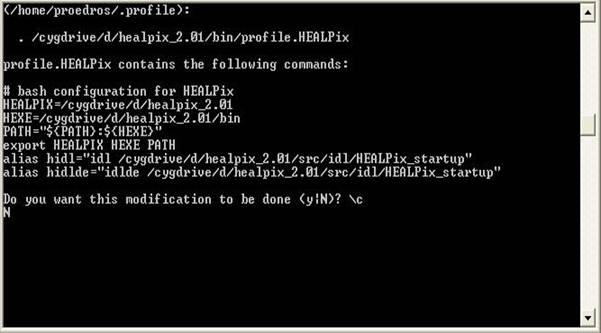
requirements section. 2) Requirements To run the codes in a Windows environment you will need: · Mathematica 5.0 + · The cygwin environment · The g95 fortran compiler · The Healpix 2.01 library · The LAPACK and BLAS libraries · The WMAP year 3 data 1a) The cygwin environment To install cygwin, go to the cygwin webpage (found here), download the cygwin setup program and run it. Follow the on screen instructions and choose only the basic packages for the time being (make sure you choose the gcc compilers and the make utility!). It will take some time to download them all as they will be ~50Mb. You need not worry about the installation as it done automatically by the executable! 1b) The g95 fortran compiler To install g95, only after you have a working cygwin enviroment, go to the g95 project webpage (found here), download the prebuilt binary appropriate for cygwin (named “self extracting Cygwin x86’’) and run it. Follow the on screen instructions and after the installation is complete run a few simple fortran programs to make sure everything is OK. To use the compiler click on the cygwin shortcut, “cd” to a directory of your choice, eg “cd c:/projects” (the “ ” are not needed) and write “g95 –o test test.f90” where the file “test.f90” is a simple Fortran program of your choice. This will create an executable named test.exe which can be run by the command: “./test”. For more options write “g95 --help”. 1c) The Healpix 2.01 library Healpix is a program that can create Temperature Anisotropy Maps from CMB data like the ones from WMAP. It is also necessary if you want to create CMB spectra in FITS format with CAMB or the likelihood of your model. To get going you will need to download the CFITSIO library and the Healpix 2.01 library. CFITSIO can be found here and you will need the Unix.tar file (cfitsio3006.tar.gz). Healpix can be found here. Once you have both, extract Healpix in a directory of your choice, say “C:/Healpix_2.01”. In the folder named “src” you can see the source files for various versions of Healpix: Fortran, C, C++ and IDL. In order to familiarize yourself with the cygwin environment, the compilers, the make utility etc, we will start with the installation of the c++ part of Healpix. Of course before you start you should at least read the documentation found in the “doc” folder in the main Healpix directory! The C++ part i. Run Cygwin and cd to the folder “C:/Healpix_2.01/src/cxx”. Notice that while Windows display the paths like this: “C:\Healpix_2.01\src\cxx” in Unix environments this is not correct. ii. Copy the cfitsio3006.tar.gz file in the “libcfitsio” folder iii. In the “Makefile” file add the line “HEALPIX_TARGET:= generic_gcc” right in the beginning and save it. Make sure this file has no extention like .txt etc. iv. Do the same with the file “rules.common” found in the folder “config” but leave the extension as it is. v. Go to the cygwin console, make sure you are in the “C:/Healpix_2.01/src/cxx” directory and type “make”. If all goes well you should have some new folders “bin”, “include” and “lib”. Now go in the “test” and type “sh ./runtest.sh”. This will run some tests and if everything is OK you should have some Temperature Anisotropy Maps! The f90 part i. Go to the main Healpix directory. Replace the file named “configure” with the one found here or add the following lines (in bold) at the end of the GuessCompiler section, so that it look like this: CYGWIN*) OFLAGS="-O" IdentifyCompiler;; *) echo "\"$OS\" is not supported yet" exit 1;; Esac ii. Go to the cygwin console, make sure you are in the “C:/Healpix_2.01” directory and type “./configure”. The script will ask you a few questions about your compiler etc. When the default answer (the one in parentheses) is the right one just hit Enter. The Session should look like this:
iii. After the configure script is done just type “make” to create the executables and the libraries iv. If you want type “make test” to test Healpix and make sure everything works out fine. For the modified version of CAMB we will only need the libraries. 1d) The LAPACK and BLAS libraries Lapack is a set of routines for matrix operations. It is necessary if you want to use the WMAP 3 likelihood routine. There two options here: The first is to download the whole Lapack and Blas packages from here and here. Detailed instructions can be found in the same pages. While Lapack is great, and is even used by Mathematica, the complete compilation can be quite big while not all of the subroutines are needed for the WMAP likelihood code, so you might want to check the second choice... The second, and easiest of the two, is to download a minimal-Lapack package from
mathfiles that contains just the appropriate routines including the ones from Blas and a makefile that compiles the library. To do so, open cygwin, go to where you extracted the package and just type “make”. Finally copy the library, “liblapack1.a”, to “C:\cygwin\lib”. 1e) The WMAP year 3 data In order to use the WMAP 3 likelihood code, found here or already integrated into the modified CAMB code supplied by the authors, obviously you will need the data. It is a rather huge download (~350Mb) but its well worth it! When the download is complete, just extract them in the directory of your choice. go to the Fortran codes section |